Welcome to the Dynamic Time Warp suite!
The packages dtw for R and dtw-python for Python provide the most complete, freely-available (GPL) implementation of Dynamic Time Warping-type (DTW) algorithms up to date. They support arbitrary local (eg symmetric, asymmetric, slope-limited) and global (windowing) constraints, fast native code, several plot styles, and more.
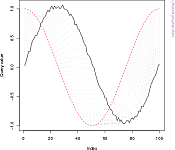
DTW is a family of algorithms which compute the local stretch or compression to apply to the time axes of two timeseries in order to optimally map one (query) onto the other (reference). DTW outputs the remaining cumulative distance between the two and, if desired, the mapping itself (warping function). DTW is widely used for classification and clustering tasks, e.g. in bioinformatics, chemometrics, econometrics, and general timeseries mining.
The R package is described in a companion paper which includes detailed instructions and extensive background on things like multivariate matching, open-end variants for real-time use, interplay between recursion types and length normalization, history, etc. The dtw-python module on PyPi is its direct Python equivalent.
Availability
- R: the dtw package on CRAN
- Python: the dtw-python package on PyPI
Both are available for all major platforms and regularly tested and built via continuous integration. Source code is available on GitHub and in the CRAN package.
Features
The implementation provides:
- arbitrary windowing functions (global constraints), eg. the Sakoe-Chiba band and the Itakura parallelogram;
- arbitrary transition types (also known as step patterns, slope
constraints, local constraints, or DP-recursion rules). This
includes dozens of well-known types:
- all step patterns classified by Rabiner-Juang, Sakoe-Chiba, and Rabiner-Myers;
- symmetric and asymmetric;
- Rabiner's smoothed variants;
- arbitrary, user-defined slope constraints
- partial matches: open-begin, open-end, substring matches
- proper, pattern-dependent, normalization (exact average distance per step)
- the Minimum Variance Matching (MVM) algorithm (Latecki et al.)
In addition to computing alignments, the package provides:
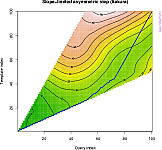
- methods for plotting alignments and warping functions in several classic styles (see plot gallery);
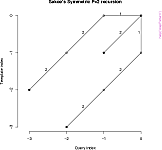
- graphical representation of step patterns;
- functions for applying a warping function, either direct or inverse;
- fast native (C) core.
Multivariate timeseries can be aligned with arbitrary local distance
definitions, leveraging the proxy::dist (R) or
scipy.spatial.distance.cdist (Python) functions.
Documentation
The best place to learn how to use the package (and a hopefully a good deal of background on DTW) is the companion paper Computing and Visualizing Dynamic Time Warping Alignments in R: The dtw Package, freely available from the Journal of Statistical Software. It includes detailed instructions and extensive background on things like multivariate matching, open-end variants for real-time use, interplay between recursion types and length normalization, history, etc.
To learn how the dtw package is used in domains ranging from bioinformatics to chemistry to data mining, please see the list of citing papers.
The R and Python pages contain links to programming language-specific documentation. (Note: R is the prime environment for the DTW suite. Python is functionally equivalent, but part of the documentation is translated automatically and may not be as pretty.)
Quickstart
Ready-to-try examples are available in the DTW for R and DTW for Python pages.
Plot gallery
See a gallery of sample plots, straight out of the examples in the documentation.
Citation
If you use dtw, do cite it in any publication reporting results
obtained with this software. Please follow the directions given in
citation("dtw"), i.e. cite:
- Toni Giorgino (2009). Computing and Visualizing Dynamic Time Warping Alignments in R: The dtw Package. Journal of Statistical Software, 31(7), 1-24, doi:10.18637/jss.v031.i07.
When using partial matching (unconstrained endpoints via the
open.begin/open.end options) and/or normalization strategies, please
also cite:
- Paolo Tormene, Toni Giorgino, Silvana Quaglini, Mario Stefanelli (2008). Matching Incomplete Time Series with Dynamic Time Warping: An Algorithm and an Application to Post-Stroke Rehabilitation. Artificial Intelligence in Medicine, 45(1), 11-34. doi:10.1016/j.artmed.2008.11.007
License
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, either version 3 of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see https://www.gnu.org/licenses/.
Contact
I am happy to provide support and seminars to academic and public research institutions. For seminars, please indicate dates, preferred format, and audience type.
Toni dot Giorgino at gmail.com
Istituto di Biofisica (IBF-CNR)
Consiglio Nazionale delle Ricerche
c/o Dept. of Biosciences, University of Milan
Milano, Italy
Commercial support
Research contracts for on-site and remote R&D for commercial companies are available through the Biophysics Institute.